JS PHOTOGRAPHY
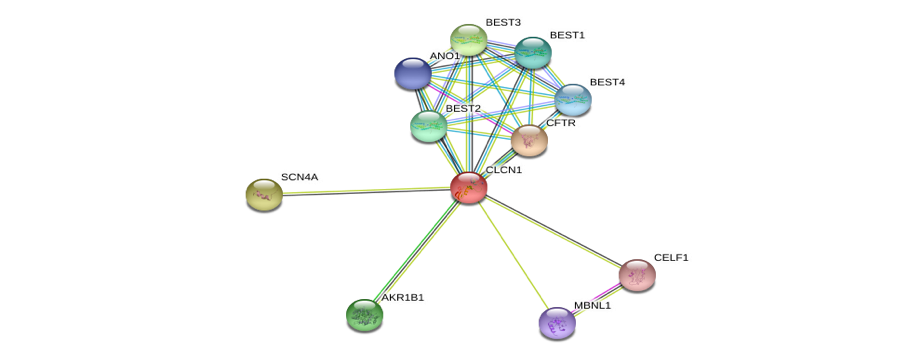
Myotonia Congenita is a genetic disorder that is characterised by the inability of skeletal muscle to relax after voluntary movement[1]. Individuals who are diagnosed with different forms of myotonia congentia typically describe a painless, muscle stiffness that remits with several repetitions of the same movement [2]. Myotonia is caused by mutations within the CLCN1 gene that decreases the chloride channel activity in the muscle in turn prolonging potential equalisation during voluntary muscle movement. The activation of chloride ion channels are essential for ensuring the electrical stability of skeletal muscle by resetting its membrane excitability to the resting state after firing an action potential [4]. There have been suggestions that Myotonia may cause infertility in males [3]. However, it is still relatively unclear how myotonia and infertility are related.
Apart from being well conserved, the mouse also has many traits which make it a great choice for a model organism. It is a vertebrate which has strong genetic similarity to humans. It also has skeletal muscle and develops quickly. Most importantly the zebrafish has a similar productive system to human. This makes the phenotypes of CLCN1 mutations easy to spot.
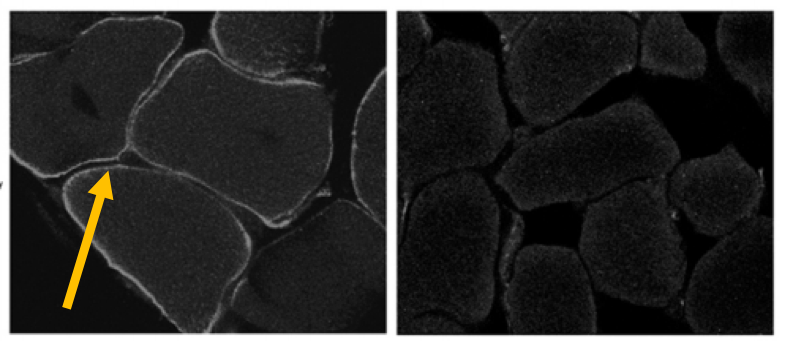
Figure 1. the figure above shows the CLCN1 protein is localized around the skeletal muscle in the wild type and the protein is not visible in the knockout.
Expression of chloride channel 1 (CLCN1/ClC-1) in skeletal muscle is driven by alternative splicing, a process regulated in part by RNA-binding protein families MBNL and CELF. Myotonia is caused by the expansion of a CTG or CCTG repeat withint thr 6B and 7A exons. This cause a increase in abnormal splicing variants or the insertion of a TAG stop codon just before exon 7 leading to a decrease in expression of the normal splice pattern.
My primary goal is to determine how the CLCN1 gene regulate chloride ions in testis. My hypothesis is that mutations in the CLCN1 gene decreases the transmembrane transport of chloride ions in turn affecting the electrochemical gradient causing sperms to be immobilised. I will use mice as my model organism because they share similar muscle structure, reproductive system, and have skeletal muscle movements that are easy to observe.
Aim 1: Determine conserved amino acids in the CLCN1 protein that mediate proper sperm function.
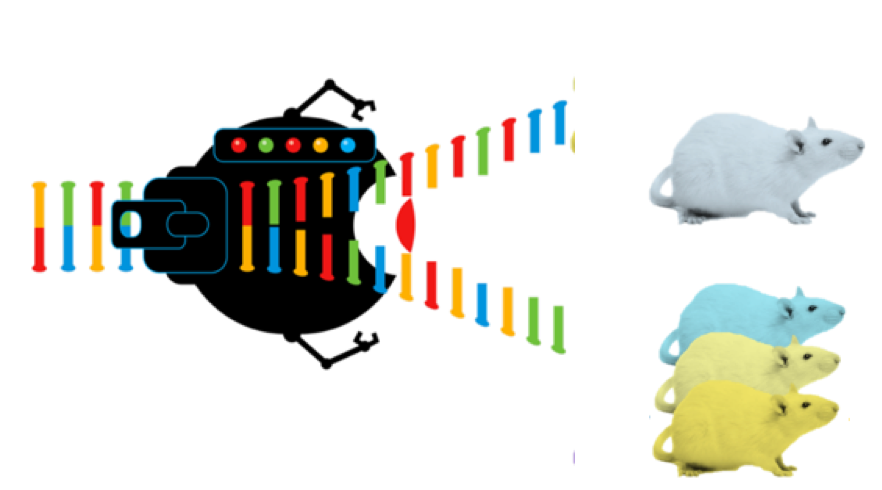
To do this, I will search for CLCN1 homologs in various species to identify conserved protein domains using the SMART database. Then, using mouse as my model organism, I will individually make targeted amino acid substitutions using CRISPR/Cas9 in the CLCN1 domain to induce myotnoia. I will do a fertility test to determine which conserved sites of CLCN1 are most critical to the proper functioning sperm.
Aim 1: Determine conserved amino acids in the CLCN1 protein that mediate proper sperm function.
To do this, I will search for CLCN1 homologs in various species to identify conserved protein domains using the SMART database. Then, using mouse as my model organism, I will individually make targeted amino acid substitutions using CRISPR/Cas9 in the CLCN1 domain to induce myotnoia. I will do a fertility test to determine which conserved sites of CLCN1 are most critical to the proper functioning sperm.
Aim 2: Determine gene expression changes necessary for fertility in CLCN1 mutant mice
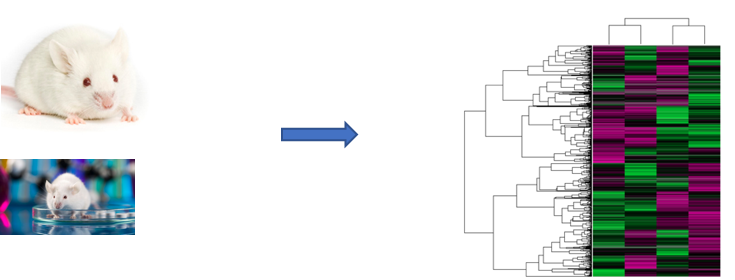
My second aim is to identify differentially expressed transcripts in the CLCN1 mutants that are important for sperm function. First, I will use RNA-seq on wild-type (WT) mice and the CLCN1 mouse mutants from Aim 1 to identify RNA transcript profiles in testis tissue and determine any differentially expressed genes. Then, through Gene Ontology (GO), I will sort expressed genes by function and identify any differences in transcript levels between the normal and disease states. I will then knock out these genes using CRISPR to determine if they show the same infertile phenotypes as the CLCN1 mutants
My second aim is to identify differentially expressed transcripts in the CLCN1 mutants that are important for sperm function. First, I will use RNA-seq on wild-type (WT) mice and the CLCN1 mouse mutants from Aim 1 to identify RNA transcript profiles in testis tissue and determine any differentially expressed genes. Then, through Gene Ontology (GO), I will sort expressed genes by function and identify any differences in transcript levels between the normal and disease states. I will then knock out these genes using CRISPR to determine if they show the same infertile phenotypes as the CLCN1 mutants
Aim 3: Identify novel CLCN1 protein interactions important for sperm development
The purpose of my final aim is to identify new protein-protein involved in sperm function. I’ll perform a Tap-Mass spectrometry on both the wild type and mutant from aim1. In addition to tap-mass spectrometry, I’ll also use a top down proteomic approach by analyzing whole tissue samples. This will also me to identify any labile post translational modifications that are lost during the digestion stage in Tap-Mass spectrometry. Finally using PANTHER, I will classify these proteins by their GO terms to determine what protein classes and biological processes are affected in the disease state.
Future directions
Overall, myotonia congentia is disease characterised by the inability of skeletal muscle to relax. It is caused by excessive CTG and CCTG nucleotide expansion within the 6th and 7th exon of the CLCN1 gene. By identifying the mechanisms that cause infertility associated with this disease will help pave way for the development of different therapies.
The purpose of my final aim is to identify new protein-protein involved in sperm function. I’ll perform a Tap-Mass spectrometry on both the wild type and mutant from aim1. In addition to tap-mass spectrometry, I’ll also use a top down proteomic approach by analyzing whole tissue samples. This will also me to identify any labile post translational modifications that are lost during the digestion stage in Tap-Mass spectrometry. Finally using PANTHER, I will classify these proteins by their GO terms to determine what protein classes and biological processes are affected in the disease state.
Future directions
Overall, myotonia congentia is disease characterised by the inability of skeletal muscle to relax. It is caused by excessive CTG and CCTG nucleotide expansion within the 6th and 7th exon of the CLCN1 gene. By identifying the mechanisms that cause infertility associated with this disease will help pave way for the development of different therapies.
| yang-5-4-18.pptx | |
| File Size: | 5862 kb |
| File Type: | pptx |
| yang-final_powerpointpptx.pdf | |
| File Size: | 4801 kb |
| File Type: | |